 An online course on single cell analysis with TranSYS ESR12.
An online course on single cell analysis with TranSYS ESR12.
One of the most exciting feelings for curious people is enlightenment on understanding how things work. Sometimes people call it “eureka”, or just say “hmm”, or “wow!” etc. And to be honest, it is the most enjoyable thing in research work.
A few years ago, I asked myself – what could be better than this? And I realized that teaching and sharing some mind-blowing concepts, which makes other people highbrow is even more pleasant.
That is why together with other scientists and bioinformaticians we started giving lectures and organizing events, where specialists in bioinformatics, data science, molecular biology, and related disciplines were able to share their expertise with other people. This is a concise story about how NGO GenomicsUA was created. Overall, we have hosted and organized more than 30 events during the last three years.
Despite the pandemic circumstances, this year was not an exception. Our team in Genomics UA has organized a one-week course of single cell analysis.
Why single-cell analysis? Single-cell sequencing is a very young and prospective technology that already has attracted thousands of scientists worldwide and received titles such as Method of the year in 2013, 2019 and 2020 from Nature journal and solved multiple enigmas in biomedicine. That is why we are so excited to share our knowledge about single-cell analysis and educate people on it.
Our school included theoretical lectures on single-cell approaches and related data analysis and hands-on training of single-cell processing. Among highlighted technologies were single-cell transcriptomics (scRNAseq), epigenomics (scATACseq), TCR/BCR sequencing, CITEseq, single-cell multiomics and much more.
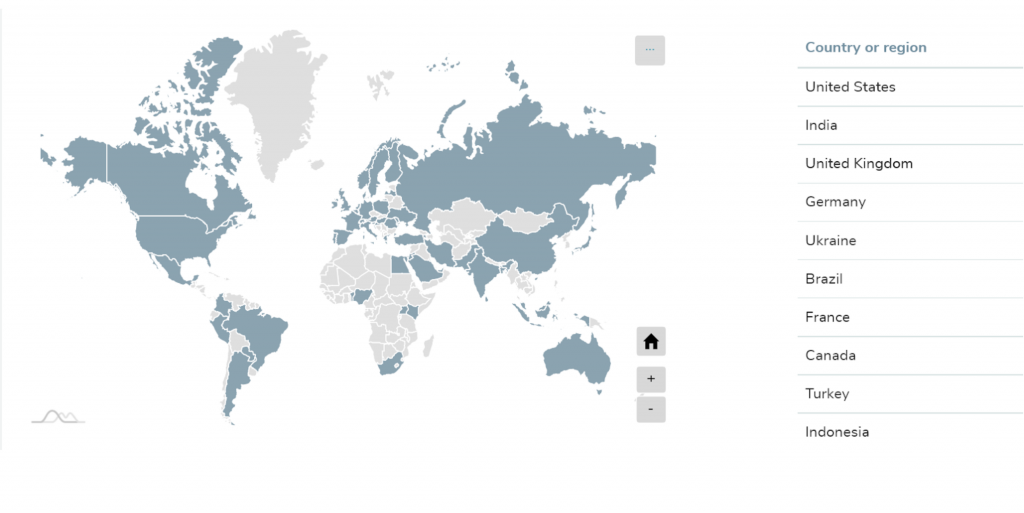
After announcing the course, we received more than 1200 applications from all parts of the world (showed on the map).
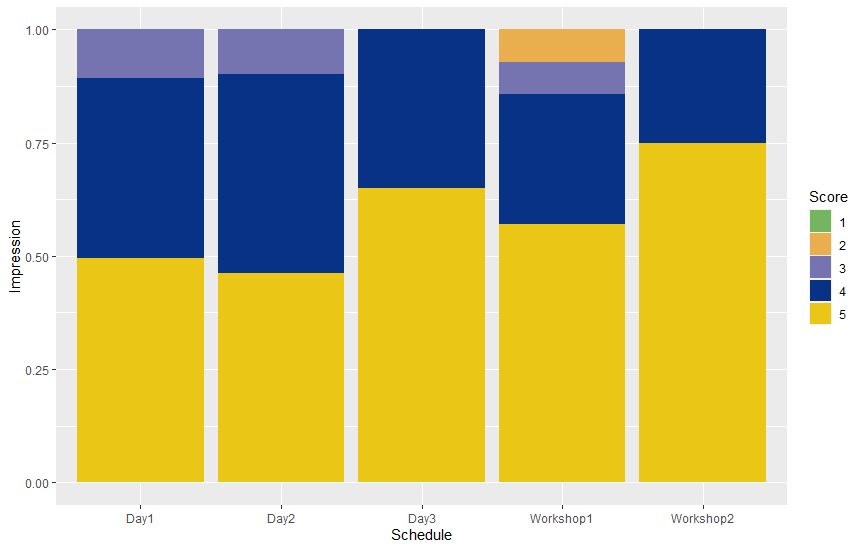
We felt a big responsibility to deliver a clear overview of single-cell technologies and help every participant to understand this field. And the feedback from our participants says that we achieved this goal! As you can see on the graph below, most participants gave us a maximum grade while evaluating the impression of the course.
 To sum up, we put a piece of our souls and did our best to run this course. It was tough but extremely enjoyable and exciting – as a lecturer and organizer, I learned a lot as well! Immediately after the closing session, our team started discussing the next course content and timeline, and we are very dedicated to organising training, schools and other events in bioinformatics in the future.
To sum up, we put a piece of our souls and did our best to run this course. It was tough but extremely enjoyable and exciting – as a lecturer and organizer, I learned a lot as well! Immediately after the closing session, our team started discussing the next course content and timeline, and we are very dedicated to organising training, schools and other events in bioinformatics in the future.
New technologies and scientific approaches appear every day. All of them are thrilling and promising, and many people would like to understand these cutting edge technologies better. However, sometimes there are some barriers to starting up with these high-level technologies. Therefore I highly recommend scientists and students to participate and organise educational events and lectures to share their awesome research and help a newbie to start with.

Author: Maryna Korshevniuk
TranSYS ESR12: Multi-omics analysis to delineate drug-response pathways
![]()

